-Search query
-Search result
Showing 1 - 50 of 52 items for (author: russell & da)
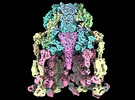
EMDB-29383: 
Structure of baseplate with receptor binding complex of Agrobacterium phage Milano
Method: single particle / : Sonani RR, Leiman PG, Wang F, Kreutzberger MAB, Sebastian A, Esteves NC, Kelly RJ, Scharf B, Egelman EH

PDB-8fqc: 
Structure of baseplate with receptor binding complex of Agrobacterium phage Milano
Method: single particle / : Sonani RR, Leiman PG, Wang F, Kreutzberger MAB, Sebastian A, Esteves NC, Kelly RJ, Scharf B, Egelman EH
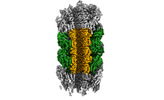
EMDB-29353: 
Structure of Agrobacterium tumefaciens bacteriophage Milano curved tail
Method: single particle / : Sonani RR, Leiman PG, Wang F, Kreutzberger MAB, Sebastian A, Esteves NC, Kelly RJ, Scharf B, Egelman EH

EMDB-29354: 
Structure of Agrobacterium tumefaciens bacteriophage Milano contracted tail-tube
Method: helical / : Sonani RR, Leiman PG, Wang F, Kreutzberger MAB, Sebastian A, Esteves NC, Kelly RJ, Scharf B, Egelman EH

EMDB-29355: 
Structure of Agrobacterium tumefaciens bacteriophage Milano contracted tail-sheath
Method: helical / : Sonani RR, Leiman PG, Wang F, Kreutzberger MAB, Sebastian A, Esteves NC, Kelly RJ, Scharf B, Egelman EH

PDB-8fop: 
Structure of Agrobacterium tumefaciens bacteriophage Milano curved tail
Method: single particle / : Sonani RR, Leiman PG, Wang F, Kreutzberger MAB, Sebastian A, Esteves NC, Kelly RJ, Scharf B, Egelman EH

PDB-8fou: 
Structure of Agrobacterium tumefaciens bacteriophage Milano contracted tail-tube
Method: helical / : Sonani RR, Leiman PG, Wang F, Kreutzberger MAB, Sebastian A, Esteves NC, Kelly RJ, Scharf B, Egelman EH

PDB-8foy: 
Structure of Agrobacterium tumefaciens bacteriophage Milano contracted tail-sheath
Method: helical / : Sonani RR, Leiman PG, Wang F, Kreutzberger MAB, Sebastian A, Esteves NC, Kelly RJ, Scharf B, Egelman EH

EMDB-27544: 
Structure of the vanadate-trapped MsbA bound to KDL
Method: single particle / : Liu C, Lyu J, Laganowsky AD, Zhao M

EMDB-27545: 
Structure of open, inward-facing MsbA from E. coli
Method: single particle / : Liu C, Lyu J, Laganowsky AD, Zhao M

PDB-8dmm: 
Structure of the vanadate-trapped MsbA bound to KDL
Method: single particle / : Liu C, Lyu J, Laganowsky AD, Zhao M

PDB-8dmo: 
Structure of open, inward-facing MsbA from E. coli
Method: single particle / : Liu C, Lyu J, Laganowsky AD, Zhao M

EMDB-26475: 
Cryo-EM structure of PAPP-A in complex with IGFBP5
Method: single particle / : Judge RA, Jain R, Hao Q, Ouch C, Sridar J, Smith CL, Wang JCK, Eaton D

EMDB-27253: 
Cryo-EM structure of substrate unbound PAPP-A
Method: single particle / : Judge RA, Jain R, Hao Q, Ouch C, Sridar J, Smith CL, Wang JCK, Eaton D

PDB-7ufg: 
Cryo-EM structure of PAPP-A in complex with IGFBP5
Method: single particle / : Judge RA, Jain R, Hao Q, Ouch C, Sridar J, Smith CL, Wang JCK, Eaton D

PDB-8d8o: 
Cryo-EM structure of substrate unbound PAPP-A
Method: single particle / : Judge RA, Jain R, Hao Q, Ouch C, Sridar J, Smith CL, Wang JCK, Eaton D

EMDB-27460: 
HMGCR-UBIAD1 Complex Dimer
Method: single particle / : Chen H, Qi X, Li X

EMDB-27461: 
HMGCR-UBIAD1 Complex Monomer
Method: single particle / : Chen H, Qi X, Li X

EMDB-27475: 
HMGCR-UBIAD1-BRIL-Fab-Nb Complex
Method: single particle / : Chen H, Qi X, Li X

EMDB-27477: 
HMGCR(40-55 deletion)-UBIAD1 Complex Dimer
Method: single particle / : Chen H, Qi X, Li X

EMDB-27478: 
HMGCR(40-55 deletion)-UBIAD1 Complex Monomer
Method: single particle / : Chen H, Qi X, Li X

EMDB-11953: 
SARS-CoV-2 S 2P trimer in complex with monovalent DARPin R2 (State 1) - Composite Map
Method: single particle / : Hurdiss DL, Drulyte I

EMDB-11954: 
SARS-CoV-2 S 2P trimer in complex with monovalent DARPin R2 (State 2)
Method: single particle / : Hurdiss DL, Drulyte I

EMDB-14810: 
SARS-CoV-2 S 2P trimer in complex with monovalent DARPin R2 (State 1) - Consensus Map
Method: single particle / : Hurdiss DL, Drulyte I

EMDB-14811: 
SARS-CoV-2 S 2P trimer in complex with monovalent DARPin R2 (State 1) - Focused Refinement
Method: single particle / : Hurdiss DL, Drulyte I

EMDB-24675: 
AMC018 SOSIP.v4.2 in complex with PGV04 Fab
Method: single particle / : Cottrell CA, de Val N, Ward AB

EMDB-24676: 
AMC016 SOSIP.v4.2 in complex with PGV04 Fab
Method: single particle / : Bader DLV, Cottrell CA, Ward AB

PDB-7rsn: 
AMC018 SOSIP.v4.2 in complex with PGV04 Fab
Method: single particle / : Cottrell CA, Ward AB

PDB-7rso: 
AMC016 SOSIP.v4.2 in complex with PGV04 Fab
Method: single particle / : Bader DLV, Cottrell CA, Ward AB

EMDB-22857: 
Structure of a ternary KRas(G13D)-SOS complex
Method: single particle / : Liu C, Moghadamchargari Z

PDB-7kfz: 
Structure of a ternary KRas(G13D)-SOS complex
Method: single particle / : Liu C, Moghadamchargari Z, Laganowsky A, Zhao M

EMDB-22295: 
Cryo-EM structure of SHIV-elicited RHA1.V2.01 in complex with HIV-1 Env BG505 DS-SOSIP.664
Method: single particle / : Gorman J, Kwong PD

PDB-6xrt: 
Cryo-EM structure of SHIV-elicited RHA1.V2.01 in complex with HIV-1 Env BG505 DS-SOSIP.664
Method: single particle / : Gorman J, Kwong PD

EMDB-7568: 
Glutaraldehyde-treated BG505 SOSIP.664 Env in complex with PGV04 Fab
Method: single particle / : Pallesen J, de Val N, Ward AB

PDB-6crq: 
Glutaraldehyde-treated BG505 SOSIP.664 Env in complex with PGV04 Fab
Method: single particle / : Pallesen J, Ward AB

EMDB-7055: 
Cryo-EM structure of the NAIP5-NLRC4-flagellin inflammasome
Method: single particle / : Tenthorey JL, Haloupek N

PDB-6b5b: 
Cryo-EM structure of the NAIP5-NLRC4-flagellin inflammasome
Method: single particle / : Tenthorey JL, Haloupek N, Lopez-Blanco JR, Grob P, Adamson E, Hartenian E, Lind NA, Bourgeois NM, Chacon P, Nogales E, Vance RE

EMDB-8699: 
EBOV GPdMuc in complex with ADI-15742 Fab
Method: single particle / : Murin CD, Ward AB, Turner HL

EMDB-6044: 
Ribosome conformation along minimum free-energy trajectory
Method: single particle / : Dashti A, Schwander P, Langlois R, Fung R, Li W, Hosseinizadeh A, Liao HY, Pallesen J, Sharma G, Stupina VA, Simon AE, Dinman J, Frank J, Ourmazd A

EMDB-2279: 
Electron cryo-microscopy of a head-tailed virus infecting extremely halophilic archaea
Method: single particle / : Pietila MK, Laurinmaki P, Russell DA, Ko CC, Jacobs-Sera D, Butcher SJ, Bamford DH, Hendrix RW

EMDB-2234: 
Electron cryo-microscopy of a head-tailed virus infecting extremely halophilic archaea
Method: single particle / : Pietila MK, Laurinmaki P, Russell DA, Ko CC, Jacobs-Sera D, Butcher SJ, Bamford DH, Hendrix RW

EMDB-2235: 
Electron cryo-microscopy of a head-tailed virus infecting extremely halophilic archaea
Method: single particle / : Pietila MK, Laurinmaki P, Russell DA, Ko CC, Jacobs-Sera D, Butcher SJ, Bamford DH, Hendrix RW

EMDB-1703: 
Intracellular part of the desmosome
Method: subtomogram averaging / : Al-Amoudi A, Frangakis AS
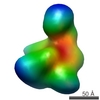
EMDB-1637: 
Proteome organization in a genome-reduced bacterium -Topoisomerase of Mycoplasma pneumoniae -
Method: single particle / : Kuhner S, vanNoort V, Betts MJ, Leo-Macias A, Batisse C, Rode M, Yamada T, Maier T, Bader S, Beltran-Alvarez P, Castano-Diez D, Chen WH, Devos D, Guell Cargol M, Norambuena T, Racke I, Rybin V, Schmidt A, Yus E, Aebersold R, Herrmann R, Bottcher B, Frangakis AS, Russell RB, Serrano L, Bork P, Gavin AC
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search








 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
